C A CBVBbc*BVBdd->C BVBda->D BVBc_->E
AND as Gene Regulation
One goal of the AND language is to capture, in simplified form, some key aspects of gene regulation:
TODO: give a statement, then detail
- Gene expression is determined by cellular context: which regulatory proteins are currently present in the cell.
- Sensitive to the DNA. The These regulatory proteins bind to specific motifs in the regulatory region upstream of the gene.
- Sensitive to local input: Other molecules, such as ligands and co-factors, influence which proteins bind and how they interact.
- Combinatorial. A combination of bound proteins determines whether transcription occurs.
- Complex networks: Gene interactions can form cascades and feedback loops: Some genes code for regulatory proteins that affect the expression of other genes.
Like other models of gene regulation, such as Boolean networks (Kauffman 1993), or thermodynamic models of transcription (Buchler et al. 2003), AND captures a few core features: Crucially, mutations to the regulatory motifs can change which proteins bind and how they interact. This means that evolutionary change can rewire gene expression without altering the proteins themselves.
Hobert (2008) models gene regulation with a similar focus on regulatory evolution. AND takes this as its central simplification: it models only regulatory evolution.
No new proteins evolve; the model focuses entirely on the conditional expression of existing genes. This follows the idea—well supported in evo-devo—that much of evolutionary change involves the reallocation and redeployment of existing functionality, not the invention of new parts (Carroll 2005).
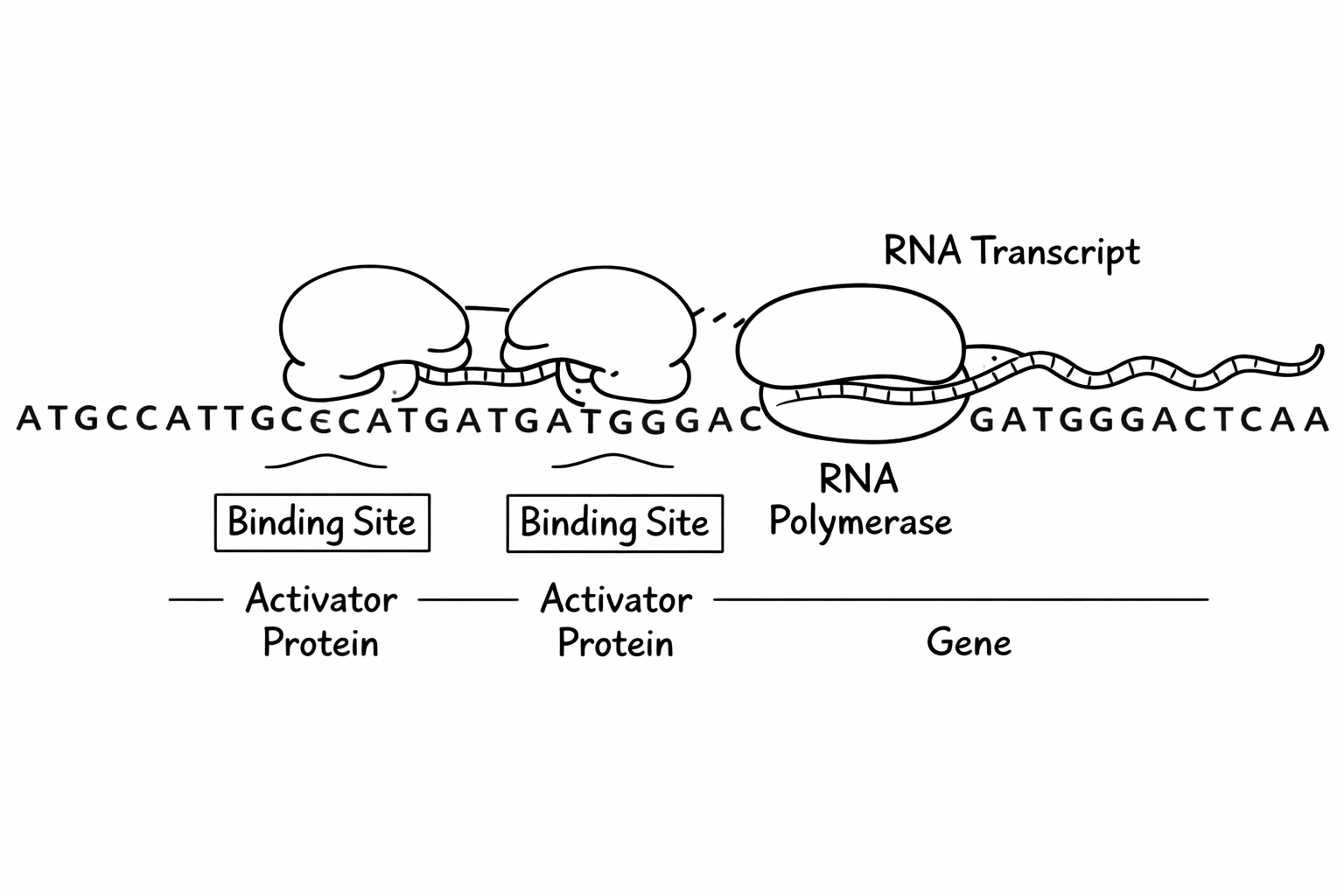
Binding and transcription in AND
With this biological picture in mind, we can map the components of an AND program onto their biological counterparts:
- The program state—the current values of all registers—represents which proteins and external factors are present in the cell.
- The syntax of each expression mimics how regulatory rules are encoded in the DNA.
- The semantics—what each expression computes—captures how the regulatory region controls gene expression.
We can treat the program from the previous section as a stretch of DNA containing regulatory regions and genes.
The structure and syntax of the modules was influenced by the discussion of “Combinatorial Transcription Logic”, in Buchler et al. (2003).
Each expression in an AND program represents a gene. The target register names the protein that gene expresses. When a gene activates, that protein will be present in the next time-step.
We distinguish gene types by which register they write to:
- An expression writing to an output register is a structural gene.
- An expression writing to a memory register is a regulatory gene.
The activation condition represents the regulatory region of the gene—encoding the conditions under which transcription occurs, and thus when the protein gets produced.
The components of the condition map onto biological terms:
- The lowercase register names are regulatory motifs that bind to whatever proteins or external factors are present:
Abinds to motifa,Bbinds to motifb, and so on. - The operators (e.g.
BVB) capture how two proteins binding at their respective motifs interact. - The combiners,
*and+, represent how multiple regulatory modules act cooperatively (*) or independently (+) to control gene expression.
The program state—the values of all registers at a given time-step—captures the current conditions in the cell.
Input registers represent conditions not directly produced by gene expression, such as ligands or co-factors. In reality, external factors influence gene expression by interacting with existing proteins; here, we simplify by allowing them to bind directly to the regulatory region.
Animation
Running the program
With the mapping in place, we can read the animation above in biological terms. Each coloured block on the timeline is a protein present in the cell at that moment. A block appearing means a gene has just been transcribed, and its product is now available to influence other genes. A block disappearing means the cell has stopped producing that protein, and—since proteins degrade—it fades from the regulatory picture. The network diagram on the right shows which regulatory regions are currently being satisfied: arrows light up when the upstream proteins are bound to the motifs that a gene’s regulatory region expects.
What the animation makes vivid is that the cell is not executing a stored plan. Nothing in the DNA says first do this, then do that. Each gene responds only to what happens to be present in the cell at the current time-step. This is the reactive character of gene regulation: expression is a continuous response to current conditions, not the step-by-step execution of a script (Harel and Pnueli 1985). The same AND program, given different stimuli, traces out different histories—because the regulatory logic is a standing disposition to respond, not a fixed sequence to be run through. When regulatory proteins feed back on their own production, the cell can also sustain states on its own, holding a pattern of expression long after the input that triggered it has gone.
Modifying a program
This is the evolutionary payoff. Because an AND expression is a regulatory region, in simplified form, a small syntactic change to that expression corresponds to a point mutation in the DNA. Change a motif from a to b and the gene now listens for a different protein. Flip an operator from BNB to BVB and a condition that required both proteins now accepts either. Swap * for + and two regulatory modules that had to cooperate now act independently. None of these changes touches the proteins themselves. The cell’s repertoire of parts is unchanged; what changes is when each part gets produced, and in response to what.
This is the kind of evolutionary move evo-devo has placed at the centre of morphological change (Carroll 2005). Most of what evolution does, on this picture, is rewire—reallocate existing genes to new times, places, and conditions—rather than invent new ones. AND makes this mechanism concrete. A single character edit can leave behaviour untouched (the motif it swaps to binds to a protein that happens never to be present), or subtly shift timing, or drop a gene out of the network altogether, or switch on a cascade that was previously silent. The same small mutational step can do nothing or do a lot, depending entirely on the regulatory context it lands in. This is what makes regulatory evolution such a productive substrate: fine-grained, open-ended, and densely connected to function, without requiring the cell to invent anything new.